SOME twelve years ago, a respected Australian geneticist predicted that the genetic potential of an animal would be measured by analysing the DNA from a tail hair or tissue sample. and that herd books and traditional Estimated Breeding Values would no longer be relevant.
It has not quite happened that way, but the introduction of Single-Step Breedplan genetic analysis in Australia is not too far from that bold prediction.
Since 2011 the Angus breed has been using a two-step genetic analysis where a standard Breedplan analysis utilising pedigrees and performance data to calculate EBVs was first run and then the results of genomic tests were blended into the EBVs of those animals with a genomic test result. This method relied heavily on the calculated correlation between the genomic test result and the trait.
Developed by Animal Genetics and Breeding Unit scientists at the University of New England, with funding from Meat & Livestock Australia, the new Single-Step approach combines traditional pedigrees, performance data and genomic results in a single analysis. It is a much more powerful method of utilising genomic information because as well as using the predictive power of the genomic test it also calculates the precise genetic relationship between all animals in the analysis.
The traditional approach assumed that 50 percent of the influential genes of a calf come from the sire and 50 percent from the dam. This is not necessarily the case, and the Single-Step approach takes account of this. The Single Step analysis also detects any incorrect parentage records on traditional pedigree databases.
The Brahman breed was the first to adopt the Single-Step analysis in April 2017, followed by the Herefords in October and Angus in December.
The Wagyu breed this week announced that it, too, is moving to a Single-Step analysis.
While it is too early for Wagyu breeders to experience the effects of the move to Single-Step, in comments below, breeders and other stakeholders from the Brahman, Hereford and Angus breeds have related their experiences with the introduction of Single-Step Breedplan analysis of their animals.
Brahmans first movers
 The Brahman breed was the first to adopt Single-Step Breedplan in April 2017 after using a two-step blending approach since 2011. The current Brahman analysis includes about 13,500 genomic test results.
The Brahman breed was the first to adopt Single-Step Breedplan in April 2017 after using a two-step blending approach since 2011. The current Brahman analysis includes about 13,500 genomic test results.
The biggest impact of utilising genomic prediction in the Brahman breed has been the significant increase in accuracy of the “Days to Calving” fertility EBVs in young bulls. This EBV is a prediction of the differences in the fertility of the future daughters of young bulls, so is a very economically important trait to the breed.
Northern Beef Technology Services technical officer, Paul Williams said there had been steady uptake of genomic testing in the Brahman breed, particularly by some of the large corporates such as Consolidated Pastoral Co and the Australian Agricultural Co, which saw the benefits of increasing the fertility of their large breeding herds, often run under challenging environmental conditions.

Paul Williams
Mr Williams said Flo Agriculture, based at Dingo, Qld, under the management of Jennifer McCamley, was also genomically testing large numbers of animals as part of an intensive selection program.
Principal of Roxborough Brahman stud in Central Queensland, Brett Coombe, said he had done a genomic test on a number of his herd sires, but does not test his sale bulls because he records a wide range of traits in Breedplan including the Days to Calving trait.
“Bull buyers look at the Days to Calving EBV on young bulls, but the increase in accuracy from the genomic test does not warrant spending $50 on a genomic test for our sale bulls,” Mr Coombe said.
“I think some Brahman breeders are using the genomic test as an alternative to performance recording, which is not the best long-term approach. If you are going to get the best value out of genomic testing, you also need to record the data on those traits,” he said.
“However I think those breeders who are just testing their better animals will gradually increase the number of animals they test, which is a positive outcome.”
Angus and Single-Step
 The first Angus analysis with the advanced Single-Step model was released in December 2017 incorporating about 35,000 genotypes.
The first Angus analysis with the advanced Single-Step model was released in December 2017 incorporating about 35,000 genotypes.
Genomic information had been incorporated in the calculation of EBVs for Angus animals since 2011 using a two-step approach where EBVs were first calculated and then the genomics results were ‘blended’ into the EBVs of animals with genomic test results.
Breed development and extension manager for Angus Australia, Andrew Byrne said the implementation of single-step analytical software into Angus BreedPlan has been welcomed by Angus breeders, particularly those that have heavily invested in genomic testing over recent years.
“Angus breeders are now benefiting from the availability of EBVs that provide a more accurate estimate of each animal’s breeding value,” he said.
“There have been noticeable changes to the EBVs for some individual animals, particularly those with genomic information, and Angus Australia has been working over recent months, in collaboration with staff at the Animal Genetics and Breeding Unit, to help members better understand the reason for these changes,” Mr Byrne said.
The most notable changes had resulted from the improved manner in which the single step analytical software models the genetic relationships between animals. Previously only pedigree information was utilised, but now both pedigree and genomic information is considered when determining how animals are related.
“This improvement has resulted in the EBVs of progeny being less aligned to the EBVs of their parents in some cases, while considerably more variation is now observed between the EBVs of full and half siblings, such as embryo transfer calves from the same flush, or progeny of the same sire,” Mr Byrne said.
“We expect that the implementation of single-step analytical software into Angus Breedplan will increase the level of genomic testing, and expect that there will be about 15,000 Angus animals genomically tested through Angus Australia during 2018,” he said.
That represents an almost 200 percent increase on testing undertaken in 2017, with the anticipated increase resulting from the availability of improved genomic products, such as the Neogen Angus GS (Genomic Selection) 50K test, and the Angus HeiferSelect commercial heifer selection tool, coupled with considerable decreases to the cost of genotyping.
Rennylea Angus at Culcairn, NSW has carried out genomic tests on its 2014 to 2017 age groups. All the retained L, M and N animals have been tested with the i50K genomics test. With some 5000 genomically tested animals, Rennylea claims to have the largest within-herd genomic database in Australian Angus.
“There have been quite large shifts in some individual bulls – some positive, some negative – but the genetics profile of the herd has not changed”
Stud principal, Bryan Corrigan, said the biggest change in performance testing since EBVs were introduced came in the December run of Angus Group Breedplan with the introduction of Single-Step.
“In the seedstock world, this has caused a lot of discussion and some negative commentary, but I think it is an exciting change and the figures will settle down and life will go on,” he said.
“Rennylea has been in a different position because around 15pc of the Angus database of 35,000 tests come from our herd. I have spent some time looking at the results in our herd. There have been quite large shifts in some individual bulls – some positive, some negative – but the genetics profile of the herd has not changed, and averages of all EBVs and indexes are very similar,” Mr Corrigan said.
“This does not surprise us. “The exciting thing is some individuals have moved significantly. We are particularly interested in how single-step is assisting us to differentiate high-value, full-flush sibs at a young age.
“We will be able to make more informed decisions particularly in relation to the young bulls we introduce into our herd and also our replacement females. These decisions are made at a young age, 400 days for the bulls and younger for some females,” he said.
“The system before didn’t differentiate between full siblings – only on individual performance. The relationship matrix clearly now shows up their correct relationship to their parents.”
Mr Corrigan said he expected the analysis would take some time to settle down, particularly in Rennylea’s case as it had so much genomic information.
“However, I am sure this will deliver more accurate EBVs which will enable clients to hone their breeding objective, which will help consistency in an emerging value-based market,” he said.

Sam White
Bald Blair Angus at Guyra, NSW has been whole-herd genomically testing for the last four years testing over 300 head each year.
Stud principal Sam White said prior to that, the stud was selective with bulls and donor dams, however it had found the whole-herd testing had led to improved selection decisions over time.
“The change to single-step has led to a refinement in the technology for the better, I think,” he said.
“I do not believe that single-step will remove the need for data collection in any way shape or form. “What we are trying to enhance is greater accuracy at a younger age with our genomically-enhanced EBV’s.”
“This is another tool which is a further refinement from the two-step analysis in the selection decision tool box,” he said.
Kilburnie Angus at Walcha, NSW has had all recent calves – 150 in 2017 and 220 in 2016 – tested with the i50K genomic test, and therefore included in the current single-step analysis.
Principal David Murray said, “We also wanted to make sure that all animals currently owned were verified to sire and dam (mating), so we have also had i50K done on all the active cows in the herd.”
“The i50K test provides the genomic data for inclusion in the single-step analysis, but for the $51.85 we currently pay for the test, we also get parent verification.”
Standalone parent verification currently costs $25.85 per test. “But cost is not insignificant. We have had at least 500 tests done at around $50 per test over the last three years, which is a total outlay of around $25,000,” Mr Murray said.
The main benefit of single-step was that the EBVs are much more precise, and Kilburnie gets the precise results without having to wait until the animal is ‘proven.’ An example is five full-sibs from the same flush born in July 2016. From the two-step analysis in mid-November 2017, the Heavy Grass Index for the five siblings ranged from +$173 to +$188. But with single-step analysis two days later, the range was $148 to +$204.
“The downside of the extra accuracy is that a couple of animals we had recently sold ‘crashed’, so we had some public relations work to do. But by the same token there were some pleasant surprises,” Mr Murray said.
“The extra accuracy of EBVs at an earlier age means we will have more confidence using young bulls and flushing maiden heifers to increase genetic progress. But we know that there is still the risk of speeding-up the rate at which we make mistakes. Not everything is in the EBVs, even the single step ones,” he said.
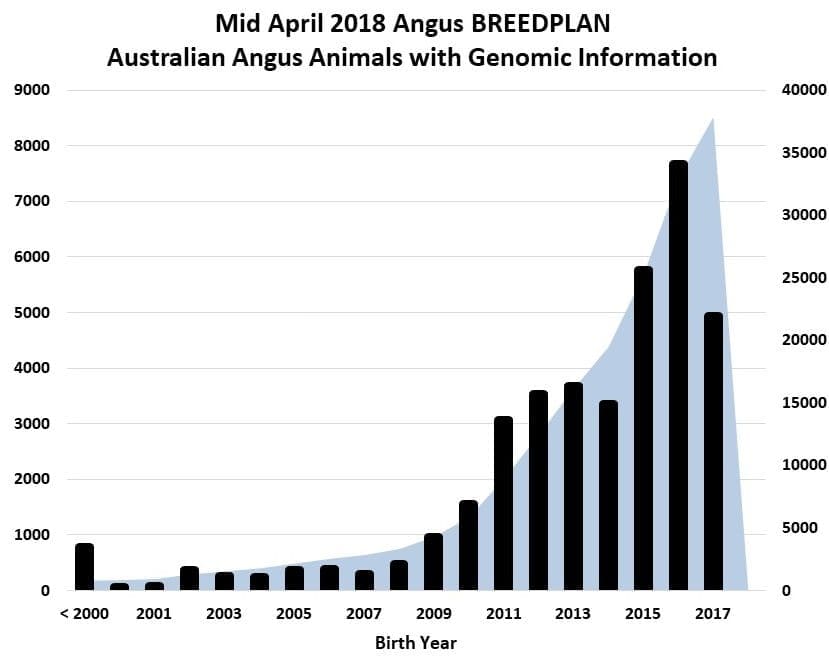
The number of Angus animals born in 2017 with a genomic test is likely to exceed the number of 2016 born animals tested, with an even bigger increase in the number of animals tested born in 2018 expected.
Herefords Australia progress
The Hereford breed undertook its first Single-Step Breedplan analysis in October 2017, after a concerted effort by former general manager Alex Ball to prepare members for the introduction of Single-Step and increase the number of genotyped animals.

Current Herefords Australia general manager Andrew Donoghue said members had been very proactive in embracing genomics and Single-Step EBVs.
“The 7086 genotypes included in the most recent analysis has grown substantially since the first Single-Step analysis, which included 3970 genotypes,” he said.
Herefords Australia was able to build an effective reference population of genotyped animals using a combination of research project animals, influential sires and other animals owned by members.
“The strategy in which the reference population was compiled led to a far quicker introduction of single-step than would have been possible, if genotyping was not so targeted,” Mr Donoghue said.
“Our current policy is that all new AI or natural sires and embryo donor cows as well as all bulls entered for the Wodonga and Dubbo National Sales (about 340 in 2018) must be genotyped. We have left it to breeders to set their own strategies for genotyping of breeding cows and young calves,” he said.
“The increased number of animals being genotyped is due to a combination of factors including the release of Single Step EBVs, members wanting to sire-verify sale bulls before sales, and the reducing cost of genotyping versus basic parent verification.”

Ian Locke
Principal of the Wirruna Hereford stud at Holbrook, Ian Locke has been one of the early-adopters of genomic testing of his 400 cow breeding herd, with more than 1000 animals genotyped over the last four years.
“I am proud that Herefords were early-adopters of the new genetic technologies and pleased that I can utilise world-best technology to capture both genomic and phenotypic information and produce more accurate genetic information that allows both my clients and I to make better breeding decisions for genetic gain in the economically important traits,” he said.
“DNA technology at Wirruna is now giving us a valuable insight into the genetics, especially carcase traits, with more accuracy and clarity than has previously been available in young bulls. Normally equivalent accuracy of information would not have been available without actually testing the progeny of these bulls,” Mr Locke said.
“In my analysis at the time of introduction, there really wasn’t any significant change in EBVs or rankings with the introduction of Single-Step in our herd. There were some small improvements in accuracy, a little bit of re-ranking of animals for some traits, but generally, nothing too significant to upset or excite,” he said.
He had found that embryo transfer flush brothers showed a greater variation in EBVs compared to what was effectively a mid-parent value previously. The EBVs of four full-flush brothers from Wirruna’s 2016 drop vary considerably, now that their genotypes have been included.
“For example their intra-muscular fat percentage EBVs range from +2.5 to +3.1 and their grainfed indexes vary from +$134 to +$158,” he said.
“A great feature of the Single-Step analysis that I appreciate is the automatic functionality to pick up incorrect parentage and or breed mix. And it even provides the correct parentage, if the correct parent has a genotype recorded.
“This may present some challenges to those who run herd books and some of their members in the initial stages, as we inevitably pick-up previous mistakes, but it is a fantastic tool because getting pedigree right is the foundation of prediction genetic performance,” Mr Locke said.
“It is still early days, and the technology is in its infancy, but I can see the analysis improving over time with more data and gene discovery.
“Genomically enhanced EBVs via Single-Step have given me more confidence to select donor cows earlier than I previously felt comfortable. This offers the opportunity to speed up genetic gain by selecting animals with more precision and using elite females earlier,” he said.
Stephen Reid, principal of the 300-cow Talbalba Hereford stud at Millmerran, QLD, said he started doing high density genomic tests on influential sires and low density genomic tests on historical and less influential sires two years ago.
“We are waiting on low-density SNPs on all of this year’s sale bulls and all retained rising three-year-old heifers after their first calf with the aim of having all cows tested in about five years,” Mr Reid said.
“We have only seen marginal movements to our EBVs since Single-Step was introduced, which surprised us as we were expecting changes. Accuracies have not changed much as yet, but may do so when the recent genomic tests are processed,” he said.
“Our cattle have a strong recording history and most recent sires have either been through the Hereford Beef Information Nucleus (BIN) or are sons of BIN sires, so perhaps this is why we have been a little underwhelmed by Single-Step so far,” Mr Reid said.
“We do believe that long term the increased linkages formed by single-step will help us with sire selection, and are therefore dedicated to going down the genomics track while continuing data collection and our reliance on Breedplan,” he said.

“Traditional” EBV’s………newer EBV’s and/or DNA trait analysis are not totally relevant AND will never be totally relevant or make other selection methods irrelevant. All are based off misguided beliefs about science and natural processes.