
AUSTRALIAN researchers have played a major role in the development of a new low-density genomics chip specifically designed for testing of tropical breeds of cattle around the world.
The genomics test is used to predict the genetic value of animals for a wide range of growth, fertility and carcase traits from a DNA sample such as hair or tissue.
The ‘GeneSeek’ Genomics Profiler-Indicus test, known locally as the GGP-TropBeef , was developed by staff at Neogen Australasia, along with Professor Ben Hayes of the University of Queensland and Dr Nick Wu from Neogen Corporation , Lincoln, Nebraska, US.
The genomic chip which has 35,000 Single Nucleotide Polymorphisms (SNPs) is suitable for a wide range of tropical breeds including Brahman, Droughtmaster, Santa Gertrudis, Brangus and tropical composites such as the Belmont Red. In addition the chip is suitable for tropical breeds more common in South America, such as the Gyr, Guzera and Nelore breeds.
Sales and marketing manager for Neogen Australasia, Sarah Buttsworth, said the GGP Tropbeef platform delivered both Bos indicus-specific SNPs for parentage testing, as well as causative assays for genetic diseases (e.g. Pompes) and production traits (e.g. polledness).
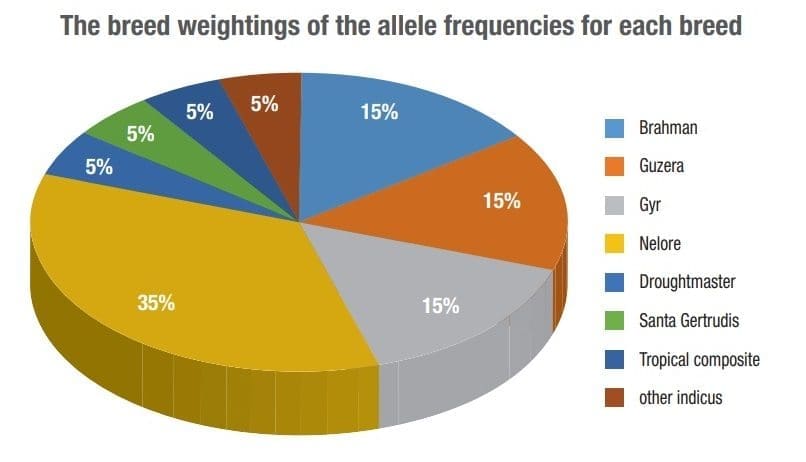
The highly-optimised 35K SNP panel included 19,000 informative backbone SNPs, 3000 SNPs specific to the Brahman and Nelore breeds, 2000 SNPs specific to the Gyr and Guzerat breeds and 3000 SNPs specific to the Droughtmaster, Santa Gertrudis and tropical composite breeds, Ms Buttsworth said.
“The advantage of a low density chip is that it is lower-priced, compared with high density chips,” she said.
“However it is important that low density chip results such as the GGP TropBeef can be imputed up to a high density genomic result with a high degree of accuracy. The accuracy of imputation of the GGP Trop Beef results to a high density (770,000 SNPs) result ranges from 94.8pc for tropical composites to 99.1pc for the Nelore breed. This is an excellent outcome,” she said.
The research population of 32,000 Brahman, Nelore, Gyr, Guzerat, Droughtmaster, Santa Gertrudis and tropical composite animals used to develop the GGP-TropBeef chip included a large number of Australian animals which were genotyped for the Co-operative Research Centre (CRC) projects, as well as the current “Repronomics” project supervised by Dr David Johnstone from the Animal Genetics and Breeding Unit at Armidale, NSW.
